Occurences as integers, e.g.ĭefault: None. The peptide mass calculator allows the determination of the mass of any peptide from its. :aa_content: A dictionary with amino acid letters as keys and it’s Parameters include molecular weight, theoretical pI, amino acid.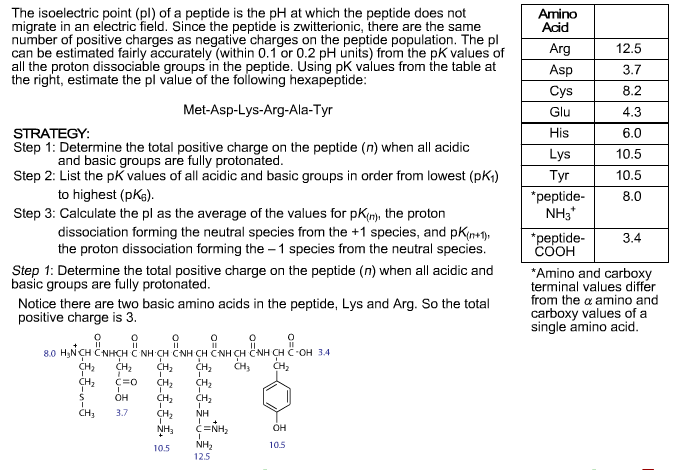 
Parameters :protein_sequence: A ``Bio.Seq`` or string object containing a protein Amino Acid Charges Biology LibreTexts Isoelectric points and amino acids (self. You will learn how to calculate the isoelectric point, and the. Online calculation (prediction) of theoretical isoelectric point (pI, IEP) of proteins and petides from sequence alone. Calculating approximate isoelectric points for El Camino A3. IsoelectricPoint ( protein_sequence, aa_content = None ) ¶Ī class for calculating the IEP or charge at given pH of a protein. The isoelectric point of an amino acid is the pH at which the amino acid has a neutral charge. For polypeptides, the pI depends primarily on the dissociation constants (Ka) of the ionizable groups of seven charged amino acids: glutamate (group -carboxyl), aspartate (group -carboxyl), cysteine (thiol group), tyrosine (phenol group), histidine (imidazole side chains), lysine (-ammonium group) and arginine (guanidinium group) plus the loa. peptide sequences are obtained by calculating the mass differences. I designed the algorithm according to a note by David L. e.g., peptide sequence, physicochemical properties (length, isoelectric point). Reference points for comparisons of two - dimensional maps of proteins from different human cell types defined in a pH scale where isoelectric points correlate with polypeptide compositions. The focusing positions of polypeptides in immobilized pH gradients can be predicted from their amino acid sequences. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
